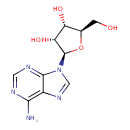
Adenosine (ECMDB00050) (M2MDB000016)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 09:56:05 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-09-13 15:15:16 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Adenosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Adenosine is nucleoside that is composed of adenine and d-ribose. Adenosine or adenosine derivatives play many important biological roles in addition to being components of DNA and RNA. Adenosine itself is a neurotransmitter. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H13N5O4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 267.2413 Monoisotopic: 267.096753929 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | OIRDTQYFTABQOQ-KQYNXXCUSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H13N5O4/c11-8-5-9(13-2-12-8)15(3-14-5)10-7(18)6(17)4(1-16)19-10/h2-4,6-7,10,16-18H,1H2,(H2,11,12,13)/t4-,6-,7-,10-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 58-61-7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2R,3R,4S,5R)-2-(6-amino-9H-purin-9-yl)-5-(hydroxymethyl)oxolane-3,4-diol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | adenosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NC1=C2N=CN([C@@H]3O[C@H](CO)[C@@H](O)[C@H]3O)C2=NC=N1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as purine nucleosides. Purine nucleosides are compounds comprising a purine base attached to a ribosyl or deoxyribosyl moiety. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Purine nucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Purine nucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 235.5 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine monophosphate + Water > Adenosine + Phosphate Adenosine + Water > Adenine + Ribose Adenosine + Adenosine triphosphate > ADP + Adenosine monophosphate + Hydrogen ion Adenosine + Hydrogen ion + Water > Inosine + Ammonium 3'-AMP + Water > Adenosine + Phosphate Adenosine + Phosphate <> Adenine + Ribose-1-phosphate Adenosine monophosphate + Water <> Adenosine + Phosphate Adenosine + Water <> Adenine + Ribose Adenosine + Water <> Inosine + Ammonia Adenosine + Phosphate <> Adenine + alpha-D-Ribose 1-phosphate Water + Adenosine > Ammonia + Inosine Adenosine + Water > D-ribose + Adenine Adenosine monophosphate + Water > Adenosine + Phosphate Adenosine + Water > beta-D-ribofuranose + Adenine Adenine + Ribose-1-phosphate > Phosphate + Adenosine Adenosine + Phosphate > Adenine + Ribose-1-phosphate Water + Adenosine > Ammonia + Inosine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Find out more about how we convert literature concentrations. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Liao, Ben-ren; Yuan, Zhen-wen. Synthesis of adenosine from inosine. Huaxue Shiji (2006), 28(10), 633-634. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in catalytic activity
- Specific function:
- A phosphate monoester + H(2)O = an alcohol + phosphate
- Gene Name:
- phoA
- Uniprot ID:
- P00634
- Molecular weight:
- 49438
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Degradation of external UDP-glucose to uridine monophosphate and glucose-1-phosphate, which can then be used by the cell
- Gene Name:
- ushA
- Uniprot ID:
- P07024
- Molecular weight:
- 60824
Reactions
| UDP-sugar + H(2)O = UMP + alpha-D-aldose 1-phosphate. |
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- This bifunctional enzyme catalyzes two consecutive reactions during ribonucleic acid degradation. Converts a 2',3'- cyclic nucleotide to a 3'-nucleotide and then the 3'-nucleotide to the corresponding nucleoside and phosphate
- Gene Name:
- cpdB
- Uniprot ID:
- P08331
- Molecular weight:
- 70832
Reactions
| Nucleoside 2',3'-cyclic phosphate + H(2)O = nucleoside 3'-phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Nucleotidase with a broad substrate specificity as it can dephosphorylate various ribo- and deoxyribonucleoside 5'- monophosphates and ribonucleoside 3'-monophosphates with highest affinity to 3'-AMP. Also hydrolyzes polyphosphate (exopolyphosphatase activity) with the preference for short-chain- length substrates (P20-25). Might be involved in the regulation of dNTP and NTP pools, and in the turnover of 3'-mononucleotides produced by numerous intracellular RNases (T1, T2, and F) during the degradation of various RNAs. Also plays a significant physiological role in stress-response and is required for the survival of E.coli in stationary growth phase
- Gene Name:
- surE
- Uniprot ID:
- P0A840
- Molecular weight:
- 26900
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| (Polyphosphate)(n) + H(2)O = (polyphosphate)(n-1) + phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows high phosphatase activity toward three nucleoside 5'-monophosphates, UMP, dUMP, and dTMP, and very low activity against TDP, IMP, UDP, GMP, dGMP, AMP, dAMP, and 6- phosphogluconate. Is strictly specific to substrates with 5'- phosphates and shows no activity against nucleoside 2'- or 3'- monophosphates. Might be involved in the pyrimidine nucleotide substrate cycles
- Gene Name:
- yjjG
- Uniprot ID:
- P0A8Y1
- Molecular weight:
- 25300
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in purine-nucleoside phosphorylase activity
- Specific function:
- Cleavage of guanosine or inosine to respective bases and sugar-1-phosphate molecules
- Gene Name:
- deoD
- Uniprot ID:
- P0ABP8
- Molecular weight:
- 25950
Reactions
| Purine nucleoside + phosphate = purine + alpha-D-ribose 1-phosphate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Uniprot ID:
- P0AE22
- Molecular weight:
- 26103
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in riboflavin synthase activity
- Specific function:
- Riboflavin synthase is a bifunctional enzyme complex catalyzing the formation of riboflavin from 5-amino-6-(1'-D)- ribityl-amino-2,4(1H,3H)-pyrimidinedione and L-3,4-dihydrohy-2- butanone-4-phosphate via 6,7-dimethyl-8-lumazine. The alpha subunit catalyzes the dismutation of 6,7-dimethyl-8-lumazine to riboflavin and 5-amino-6-(1'-D)-ribityl-amino-2,4(1H,3H)- pyrimidinedione
- Gene Name:
- ribE
- Uniprot ID:
- P0AFU8
- Molecular weight:
- 23445
Reactions
| 2 6,7-dimethyl-8-(1-D-ribityl)lumazine = riboflavin + 4-(1-D-ribitylamino)-5-amino-2,6-dihydroxypyrimidine. |
- General function:
- Involved in deaminase activity
- Specific function:
- Adenosine + H(2)O = inosine + NH(3)
- Gene Name:
- add
- Uniprot ID:
- P22333
- Molecular weight:
- 36397
Reactions
| Adenosine + H(2)O = inosine + NH(3). |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes both purine and pyrimidine ribonucleosides with a broad-substrate specificity with decreasing activity in the order uridine, xanthosine, inosine, adenosine, cytidine, guanosine
- Gene Name:
- rihC
- Uniprot ID:
- P22564
- Molecular weight:
- 32560
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes cytidine or uridine to ribose and cytosine or uracil, respectively. Has a clear preference for cytidine over uridine. Strictly specific for ribonucleosides. Has a low but significant activity for the purine nucleoside xanthosine
- Gene Name:
- rihB
- Uniprot ID:
- P33022
- Molecular weight:
- 33748
Reactions
| A pyrimidine nucleoside + H(2)O = D-ribose + a pyrimidine base. |
- General function:
- Involved in ATP binding
- Specific function:
- Catalyzes the reversible transfer of the terminal phosphate group between ATP and AMP. This small ubiquitous enzyme involved in the energy metabolism and nucleotide synthesis, is essential for maintenance and cell growth
- Gene Name:
- adk
- Uniprot ID:
- P69441
- Molecular weight:
- 23586
Reactions
| ATP + AMP = 2 ADP. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows strict specificity toward deoxyribonucleoside 5'-monophosphates and does not dephosphorylate 5'-ribonucleotides or ribonucleoside 3'-monophosphates. Might be involved in the regulation of all dNTP pools in E.coli
- Gene Name:
- yfbR
- Uniprot ID:
- P76491
- Molecular weight:
- 22708
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Energy production and conversion
- Specific function:
- Deaminates adenosine-34 to inosine in tRNA-Arg2. Mutation in this protein makes E.coli resistant to the toxic proteins encoded by the gef gene family. Essential for cell viability
- Gene Name:
- tadA
- Uniprot ID:
- P68398
- Molecular weight:
- 18717
Reactions
| Adenosine + H(2)O = inosine + NH(3). |
Transporters
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Nucleoside transport
- Gene Name:
- xapB
- Uniprot ID:
- P45562
- Molecular weight:
- 46139
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Transports nucleosides with a high affinity except guanosine and deoxyguanosine. Driven by a proton motive force
- Gene Name:
- nupC
- Uniprot ID:
- P0AFF2
- Molecular weight:
- 43475
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Transports nucleosides with a high affinity. Driven by a proton motive force
- Gene Name:
- nupG
- Uniprot ID:
- P0AFF4
- Molecular weight:
- 46389
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Nucleoside transporter
- Gene Name:
- nupX
- Uniprot ID:
- P33021
- Molecular weight:
- 43409
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Constitutes the receptor for colicin K and phage T6, and functions as substrate-specific channel for nucleosides and deoxynucleosides
- Gene Name:
- tsx
- Uniprot ID:
- P0A927
- Molecular weight:
- 33589