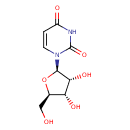
Uridine (ECMDB00296) (M2MDB000124)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 10:25:56 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-09-13 12:56:07 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Uridine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Uridine is a molecule (known as a nucleoside) that is formed when uracil is attached to a ribose ring (also known as a ribofuranose) via a b-N1-glycosidic bond. (Wikipedia) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C9H12N2O6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 244.2014 Monoisotopic: 244.069536126 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | DRTQHJPVMGBUCF-XVFCMESISA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C9H12N2O6/c12-3-4-6(14)7(15)8(17-4)11-2-1-5(13)10-9(11)16/h1-2,4,6-8,12,14-15H,3H2,(H,10,13,16)/t4-,6-,7-,8-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 58-96-8 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-1,2,3,4-tetrahydropyrimidine-2,4-dione | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | uridine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)N1C=CC(=O)NC1=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as pyrimidine nucleosides. Pyrimidine nucleosides are compounds comprising a pyrimidine base attached to a ribosyl or deoxyribosyl moiety. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pyrimidine nucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Pyrimidine nucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 163 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Water + Uridine 5'-monophosphate > Phosphate + Uridine Water + Uridine > Ribose + Uracil Guanosine triphosphate + Uridine > Guanosine diphosphate + Hydrogen ion + Uridine 5'-monophosphate Cytidine + Hydrogen ion + Water > Ammonium + Uridine Phosphate + Uridine <> Ribose-1-phosphate + Uracil 3'-UMP + Water > Phosphate + Uridine Uridine 5'-monophosphate + Water <> Uridine + Phosphate Adenosine triphosphate + Uridine <> ADP + Uridine 5'-monophosphate Uridine triphosphate + Uridine <> Uridine 5'-diphosphate + Uridine 5'-monophosphate Guanosine triphosphate + Uridine <> Guanosine diphosphate + Uridine 5'-monophosphate Inosine triphosphate + Uridine <> IDP + Uridine 5'-monophosphate dATP + Uridine <> dADP + Uridine 5'-monophosphate Uridine + Phosphate <> Uracil + alpha-D-Ribose 1-phosphate + Ribose-1-phosphate Cytidine + Water <> Uridine + Ammonia dGTP + Uridine <> dGDP + Uridine 5'-monophosphate Thymidine 5'-triphosphate + Uridine <> dTDP + Uridine 5'-monophosphate dCTP + Uridine <> dCDP + Uridine 5'-monophosphate Deoxyuridine triphosphate + Uridine <> dUDP + Uridine 5'-monophosphate Water + Cytidine > Ammonia + Uridine Uridine + Water > D-ribose + Uracil Uridine + Inorganic phosphate > Uracil + Ribose-1-phosphate Adenosine triphosphate + Uridine > ADP + Uridine 5'-monophosphate Cytidine + Water + Deoxycytidine <> Uridine + Ammonia + Deoxyuridine Uridine + Phosphate > Uracil + Ribose-1-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Find out more about how we convert literature concentrations. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Doi, Muneharu; Asahi, Satoru; Izawa, Motoo. Fermentative production of uridine and cytidine. Baiosaiensu to Indasutori (1993), 51(12), 972-6. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in hydrolase activity
- Specific function:
- Degradation of external UDP-glucose to uridine monophosphate and glucose-1-phosphate, which can then be used by the cell
- Gene Name:
- ushA
- Uniprot ID:
- P07024
- Molecular weight:
- 60824
Reactions
| UDP-sugar + H(2)O = UMP + alpha-D-aldose 1-phosphate. |
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- This bifunctional enzyme catalyzes two consecutive reactions during ribonucleic acid degradation. Converts a 2',3'- cyclic nucleotide to a 3'-nucleotide and then the 3'-nucleotide to the corresponding nucleoside and phosphate
- Gene Name:
- cpdB
- Uniprot ID:
- P08331
- Molecular weight:
- 70832
Reactions
| Nucleoside 2',3'-cyclic phosphate + H(2)O = nucleoside 3'-phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Nucleotidase with a broad substrate specificity as it can dephosphorylate various ribo- and deoxyribonucleoside 5'- monophosphates and ribonucleoside 3'-monophosphates with highest affinity to 3'-AMP. Also hydrolyzes polyphosphate (exopolyphosphatase activity) with the preference for short-chain- length substrates (P20-25). Might be involved in the regulation of dNTP and NTP pools, and in the turnover of 3'-mononucleotides produced by numerous intracellular RNases (T1, T2, and F) during the degradation of various RNAs. Also plays a significant physiological role in stress-response and is required for the survival of E.coli in stationary growth phase
- Gene Name:
- surE
- Uniprot ID:
- P0A840
- Molecular weight:
- 26900
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| (Polyphosphate)(n) + H(2)O = (polyphosphate)(n-1) + phosphate. |
- General function:
- Involved in ATP binding
- Specific function:
- ATP + uridine = ADP + UMP
- Gene Name:
- udk
- Uniprot ID:
- P0A8F4
- Molecular weight:
- 24353
Reactions
| ATP + uridine = ADP + UMP. |
| ATP + cytidine = ADP + CMP. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows high phosphatase activity toward three nucleoside 5'-monophosphates, UMP, dUMP, and dTMP, and very low activity against TDP, IMP, UDP, GMP, dGMP, AMP, dAMP, and 6- phosphogluconate. Is strictly specific to substrates with 5'- phosphates and shows no activity against nucleoside 2'- or 3'- monophosphates. Might be involved in the pyrimidine nucleotide substrate cycles
- Gene Name:
- yjjG
- Uniprot ID:
- P0A8Y1
- Molecular weight:
- 25300
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in zinc ion binding
- Specific function:
- This enzyme scavenge exogenous and endogenous cytidine and 2'-deoxycytidine for UMP synthesis
- Gene Name:
- cdd
- Uniprot ID:
- P0ABF6
- Molecular weight:
- 31539
Reactions
| Cytidine + H(2)O = uridine + NH(3). |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Uniprot ID:
- P0AE22
- Molecular weight:
- 26103
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in transferase activity, transferring pentosyl groups
- Specific function:
- Catalyzes the reversible phosphorylytic cleavage of uridine and deoxyuridine to uracil and ribose- or deoxyribose-1- phosphate. The produced molecules are then utilized as carbon and energy sources or in the rescue of pyrimidine bases for nucleotide synthesis
- Gene Name:
- udp
- Uniprot ID:
- P12758
- Molecular weight:
- 27159
Reactions
| Uridine + phosphate = uracil + alpha-D-ribose 1-phosphate. |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes both purine and pyrimidine ribonucleosides with a broad-substrate specificity with decreasing activity in the order uridine, xanthosine, inosine, adenosine, cytidine, guanosine
- Gene Name:
- rihC
- Uniprot ID:
- P22564
- Molecular weight:
- 32560
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes cytidine or uridine to ribose and cytosine or uracil, respectively. Has a clear preference for cytidine over uridine. Strictly specific for ribonucleosides. Has a low but significant activity for the purine nucleoside xanthosine
- Gene Name:
- rihB
- Uniprot ID:
- P33022
- Molecular weight:
- 33748
Reactions
| A pyrimidine nucleoside + H(2)O = D-ribose + a pyrimidine base. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows strict specificity toward deoxyribonucleoside 5'-monophosphates and does not dephosphorylate 5'-ribonucleotides or ribonucleoside 3'-monophosphates. Might be involved in the regulation of all dNTP pools in E.coli
- Gene Name:
- yfbR
- Uniprot ID:
- P76491
- Molecular weight:
- 22708
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes with equal efficiency cytidine or uridine to ribose and cytosine or uracil, respectively
- Gene Name:
- rihA
- Uniprot ID:
- P41409
- Molecular weight:
- 33823
Transporters
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Nucleoside transport
- Gene Name:
- xapB
- Uniprot ID:
- P45562
- Molecular weight:
- 46139
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Transports nucleosides with a high affinity except guanosine and deoxyguanosine. Driven by a proton motive force
- Gene Name:
- nupC
- Uniprot ID:
- P0AFF2
- Molecular weight:
- 43475
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Transports nucleosides with a high affinity. Driven by a proton motive force
- Gene Name:
- nupG
- Uniprot ID:
- P0AFF4
- Molecular weight:
- 46389
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Nucleoside transporter
- Gene Name:
- nupX
- Uniprot ID:
- P33021
- Molecular weight:
- 43409
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Constitutes the receptor for colicin K and phage T6, and functions as substrate-specific channel for nucleosides and deoxynucleosides
- Gene Name:
- tsx
- Uniprot ID:
- P0A927
- Molecular weight:
- 33589