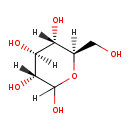
D-Glucose (ECMDB00122) (M2MDB000045)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 09:57:45 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-06-03 15:53:12 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | D-Glucose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Glucose is a monosaccharide containing six carbon atoms and an aldehyde group and is therefore referred to as an aldohexose. The glucose molecule can exist in an open-chain (acyclic) and ring (cyclic) form, the latter being the result of an intramolecular reaction between the aldehyde C atom and the C-5 hydroxyl group to form an intramolecular hemiacetal. In water solution both forms are in equilibrium and at pH 7 the cyclic one is the predominant. Glucose is a primary source of energy for living organisms. It is naturally occurring and is found in fruits and other parts of plants in its free state. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C6H12O6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 180.1559 Monoisotopic: 180.063388116 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | WQZGKKKJIJFFOK-GASJEMHNSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C6H12O6/c7-1-2-3(8)4(9)5(10)6(11)12-2/h2-11H,1H2/t2-,3-,4+,5-,6?/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 50-99-7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (3R,4S,5S,6R)-6-(hydroxymethyl)oxane-2,3,4,5-tetrol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | glucose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | OC[C@H]1OC(O)[C@H](O)[C@@H](O)[C@@H]1O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as hexoses. These are monosaccharides in which the sugar unit is a is a six-carbon containing moeity. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic oxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Carbohydrates and carbohydrate conjugates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Hexoses | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 146-150 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Phosphoenolpyruvic acid + D-Glucose > Glucose 6-phosphate + Pyruvic acid Ubiquinone-8 + D-Glucose + Water > Ubiquinol-8 + Gluconic acid + Hydrogen ion Adenosine triphosphate + Water + D-Glucose > ADP + D-Glucose + Hydrogen ion + Phosphate Adenosine triphosphate + Water + D-Glucose > ADP + D-Glucose + Hydrogen ion + Phosphate Water + alpha-Lactose > D-Galactose + D-Glucose Water + Maltotriose > D-Glucose + D-Maltose Water + Maltotetraose > D-Glucose + Maltotriose Water + Maltoheptaose > D-Glucose + Maltohexaose Water + Maltohexaose > D-Glucose + Maltopentaose Water + Maltopentaose > D-Glucose + Maltotetraose Acetyl-CoA + D-Glucose <> 6-Acetyl-D-glucose + Coenzyme A Glucose 6-phosphate + Water > D-Glucose + Phosphate Glucose 1-phosphate + Water > D-Glucose + Phosphate Water + Trehalose >2 D-Glucose Adenosine triphosphate + D-Glucose > ADP + Glucose 6-phosphate + Hydrogen ion D-Maltose + Maltotriose > D-Glucose + Maltotetraose D-Maltose + Maltotetraose > D-Glucose + Maltopentaose D-Maltose + Maltohexaose > D-Glucose + Maltoheptaose D-Maltose + Maltopentaose > D-Glucose + Maltohexaose D-Glucose <> D-Fructose Water + Melibiose > D-Galactose + D-Glucose Water + Trehalose 6-phosphate > Glucose 6-phosphate + D-Glucose Trehalose + Water <>2 D-Glucose Adenosine triphosphate + D-Glucose <> ADP + D-Hexose 6-phosphate + Glucose 6-phosphate Sucrose + Water <> D-Fructose + D-Glucose Water + Trehalose 6-phosphate <> D-Glucose + D-Hexose 6-phosphate More...Melibiose + Water <> D-Galactose + D-Glucose Protein N(pi)-phospho-L-histidine + D-Glucose <> Protein histidine + Glucose 6-phosphate Cyanoglycoside + Water <> Cyanohydrin + D-Glucose cis-beta-D-Glucosyl-2-hydroxycinnamate + Water <> cis-2-Hydroxycinnamate + D-Glucose 1,4-alpha-D-glucan + D-Glucose <> D-Maltose Neohancoside D + Water <> D-Fructose + D-Glucose Melibiose + Water <> D-Galactose + D-Glucose Water + Trehalose 6-phosphate <> D-Glucose + D-Hexose 6-phosphate Lactose + Water <> D-Glucose + D-Galactose D-Glucose + Ubiquinone-1 <> Gluconolactone + Ubiquinol-8 D-Glucose <> b-D-Glucose a 1,4-α-D-glucan + Water > a 1,4-α-D-glucan + D-Glucose Maltotriose + Water > D-Maltose + D-Glucose nigerose + Water D-Glucose Trehalose + Water > b-D-Glucose + D-Glucose Alpha-D-glucose 1-phosphate + Water > D-Glucose + Inorganic phosphate 6-Phospho-beta-D-glucosyl-(1,4)-D-glucose + Water > D-Glucose + Glucose 6-phosphate D-Glucose + Ubiquinone-10 > Gluconolactone + Ubiquinol-1 Adenosine triphosphate + D-Glucose > ADP + Glucose 6-phosphate Cellobiose-6-phosphate + Water <> D-Glucose + Glucose 6-phosphate Glucose 1-phosphate + Water <> D-Glucose + Phosphate Trehalose 6-phosphate + Water <> D-Glucose + Glucose 6-phosphate D-Glucose + Adenosine triphosphate > Hydrogen ion + Adenosine diphosphate + beta-D-Glucose 6-phosphate + ADP Sucrose + Water <> D-Fructose + D-Glucose + D-Fructose D-Glucose + [PTS enzyme I]-Nπ-phospho-L-histidine > β-D-glucose 1-phosphate + [PTS enzyme I]-L-histidine D-Glucose + Ubiquinone-1 <> Gluconolactone + Ubiquinol-8 Water + Trehalose >2 D-Glucose D-Glucose + Ubiquinone-1 <> Gluconolactone + Ubiquinol-8 Trehalose + Water <>2 D-Glucose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Li, Dalin; Ruan, Yi; Song, Wen; Wang, Yongjun. Improved process for producing glucose. Faming Zhuanli Shenqing Gongkai Shuomingshu (2003), 4 pp | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in beta-galactosidase activity
- Specific function:
- Hydrolysis of terminal non-reducing beta-D- galactose residues in beta-D-galactosides
- Gene Name:
- lacZ
- Uniprot ID:
- P00722
- Molecular weight:
- 116482
Reactions
| Hydrolysis of terminal non-reducing beta-D-galactose residues in beta-D-galactosides. |
- General function:
- Involved in xylose isomerase activity
- Specific function:
- D-xylose = D-xylulose
- Gene Name:
- xylA
- Uniprot ID:
- P00944
- Molecular weight:
- 49742
Reactions
| D-xylose = D-xylulose. |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- Hydrolysis of terminal, non-reducing alpha-D- galactose residues in alpha-D-galactosides, including galactose oligosaccharides, galactomannans and galactolipids
- Gene Name:
- melA
- Uniprot ID:
- P06720
- Molecular weight:
- 50657
Reactions
| Hydrolysis of terminal, non-reducing alpha-D-galactose residues in alpha-D-galactosides, including galactose oligosaccharides, galactomannans and galactolipids. |
- General function:
- Involved in beta-galactosidase activity
- Specific function:
- The wild-type enzyme is an ineffective lactase. Two classes of point mutations dramatically improve activity of the enzyme
- Gene Name:
- ebgA
- Uniprot ID:
- P06864
- Molecular weight:
- 117878
Reactions
| Hydrolysis of terminal non-reducing beta-D-galactose residues in beta-D-galactosides. |
- General function:
- Involved in transferase activity, transferring phosphorus-containing groups
- Specific function:
- General (non sugar-specific) component of the phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS). This major carbohydrate active-transport system catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. Enzyme I transfers the phosphoryl group from phosphoenolpyruvate (PEP) to the phosphoryl carrier protein (HPr)
- Gene Name:
- ptsI
- Uniprot ID:
- P08839
- Molecular weight:
- 63561
Reactions
| Phosphoenolpyruvate + protein L-histidine = pyruvate + protein N(pi)-phospho-L-histidine. |
- General function:
- Involved in glucokinase activity
- Specific function:
- Not highly important in E.coli as glucose is transported into the cell by the PTS system already as glucose 6-phosphate
- Gene Name:
- glk
- Uniprot ID:
- P0A6V8
- Molecular weight:
- 34723
Reactions
| ATP + D-glucose = ADP + D-glucose 6-phosphate. |
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex MglABC involved in galactose/methyl galactoside import. Responsible for energy coupling to the transport system (Probable)
- Gene Name:
- mglA
- Uniprot ID:
- P0AAG8
- Molecular weight:
- 56415
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- Can hydrolyze salicin and arbutin
- Gene Name:
- bglB
- Uniprot ID:
- P11988
- Molecular weight:
- 53161
Reactions
| 6-phospho-beta-D-glucosyl-(1,4)-D-glucose + H(2)O = D-glucose + D-glucose 6-phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Provides the cells with the ability to utilize trehalose at high osmolarity by splitting it into glucose molecules that can subsequently be taken up by the phosphotransferase-mediated uptake system
- Gene Name:
- treA
- Uniprot ID:
- P13482
- Molecular weight:
- 63636
Reactions
| Alpha,alpha-trehalose + H(2)O = 2 D-glucose. |
- General function:
- Involved in oxidoreductase activity, acting on CH-OH group of donors
- Specific function:
- GDH is probably involved in energy conservation rather than in sugar metabolism
- Gene Name:
- gcd
- Uniprot ID:
- P15877
- Molecular weight:
- 86747
Reactions
| D-glucose + ubiquinone = D-glucono-1,5-lactone + ubiquinol. |
- General function:
- Involved in 4-alpha-glucanotransferase activity
- Specific function:
- Transfers a segment of a (1->4)-alpha-D-glucan to a new position in an acceptor, which may be glucose or a (1->4)-alpha-D-glucan
- Gene Name:
- malQ
- Uniprot ID:
- P15977
- Molecular weight:
- 78503
Reactions
| Transfers a segment of a (1->4)-alpha-D-glucan to a new position in an acceptor, which may be glucose or a (1->4)-alpha-D-glucan. |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- Hydrolyzes a wide variety of P-beta-glucosides including cellobiose-6P, salicin-6P, arbutin-6P, gentiobiose-6P, methyl- beta-glucoside-6P and p-nitrophenyl-beta-D-glucopyranoside-6P. Is also able to hydrolyze phospho-N,N'-diacetylchitobiose
- Gene Name:
- chbF
- Uniprot ID:
- P17411
- Molecular weight:
- 50512
Reactions
| 6-phospho-beta-D-glucosyl-(1,4)-D-glucose + H(2)O = D-glucose + D-glucose 6-phosphate. |
- General function:
- Involved in protein-N(PI)-phosphohistidine-sugar phosphotransferase activity
- Specific function:
- MalX encodes a phosphotransferase system enzyme II that can recognize glucose and maltose as substrates even though these sugars may not represent the natural substrates of the system
- Gene Name:
- malX
- Uniprot ID:
- P19642
- Molecular weight:
- 56627
Reactions
| Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Absolutely required for the growth of E.coli in a high- phosphate medium containing G-1-P as the sole carbon source
- Gene Name:
- agp
- Uniprot ID:
- P19926
- Molecular weight:
- 45683
Reactions
| Alpha-D-glucose 1-phosphate + H(2)O = D-glucose + phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- May play a role in regulating the intracellular level of maltotriose. Cleaves glucose from the reducing end of maltotriose and longer maltodextrins with a chain length of up to 7 glucose units
- Gene Name:
- malZ
- Uniprot ID:
- P21517
- Molecular weight:
- 69172
Reactions
| Hydrolysis of terminal, non-reducing (1->4)-linked alpha-D-glucose residues with release of alpha-D-glucose. |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- Can hydrolyze salicin, cellobiose, and probably arbutin
- Gene Name:
- ascB
- Uniprot ID:
- P24240
- Molecular weight:
- 53935
Reactions
| 6-phospho-beta-D-glucosyl-(1,4)-D-glucose + H(2)O = D-glucose + D-glucose 6-phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Alpha,alpha-trehalose 6-phosphate + H(2)O = D- glucose + D-glucose 6-phosphate
- Gene Name:
- treC
- Uniprot ID:
- P28904
- Molecular weight:
- 63837
Reactions
| Alpha,alpha-trehalose 6-phosphate + H(2)O = D-glucose + D-glucose 6-phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Exhibits hydrolysis activity against alpha-glucosyl fluoride, although natural substrates, such as alpha-glucobioses are scarcely hydrolyzed
- Gene Name:
- yihQ
- Uniprot ID:
- P32138
- Molecular weight:
- 77274
Reactions
| Hydrolysis of terminal, non-reducing (1->4)-linked alpha-D-glucose residues with release of alpha-D-glucose. |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- Hydrolysis of terminal, non-reducing beta-D- glucosyl residues with release of beta-D-glucose
- Gene Name:
- bglX
- Uniprot ID:
- P33363
- Molecular weight:
- 83459
Reactions
| Hydrolysis of terminal, non-reducing beta-D-glucosyl residues with release of beta-D-glucose. |
- General function:
- Involved in catalytic activity
- Specific function:
- Hydrolyzes trehalose to glucose. Could be involved, in cells returning to low osmolarity conditions, in the utilization of the accumulated cytoplasmic trehalose, which was synthesized in response to high osmolarity
- Gene Name:
- treF
- Uniprot ID:
- P62601
- Molecular weight:
- 63696
Reactions
| Alpha,alpha-trehalose + H(2)O = 2 D-glucose. |
- General function:
- Involved in sugar:hydrogen symporter activity
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. This system is involved in glucose transport
- Gene Name:
- crr
- Uniprot ID:
- P69783
- Molecular weight:
- 18251
Reactions
| Protein EIIA N(pi)-phospho-L-histidine + protein EIIB = protein EIIA + protein EIIB N(pi)-phospho-L-histidine/cysteine. |
- General function:
- Involved in protein-N(PI)-phosphohistidine-sugar phosphotransferase activity
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. This system is involved in glucose transport. This enzyme is also a chemoreceptor monitoring the environment for changes in sugar concentration
- Gene Name:
- ptsG
- Uniprot ID:
- P69786
- Molecular weight:
- 50676
Reactions
| Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate. |
- General function:
- Involved in phosphoenolpyruvate-dependent sugar phosphotransferase system
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. This system is involved in mannose transport
- Gene Name:
- manX
- Uniprot ID:
- P69797
- Molecular weight:
- 35047
Reactions
| Protein EIIA N(pi)-phospho-L-histidine + protein EIIB = protein EIIA + protein EIIB N(pi)-phospho-L-histidine/cysteine. |
| Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the hydrolysis of sugar phosphate to sugar and inorganic phosphate. Has a wide substrate specificity catalyzing the hydrolysis of fructose-1-P most efficiently, but it remains uncertain if this is the real substrate in vivo
- Gene Name:
- supH
- Uniprot ID:
- P75792
- Molecular weight:
- 30413
Reactions
| Sugar phosphate + H(2)O = sugar + phosphate. |
- General function:
- Involved in transferase activity
- Specific function:
- Acetylates maltose and other sugars
- Gene Name:
- maa
- Uniprot ID:
- P77791
- Molecular weight:
- 20096
Reactions
| Acetyl-CoA + maltose = CoA + acetyl-maltose. |
- General function:
- Involved in hydrolase activity, hydrolyzing O-glycosyl compounds
- Specific function:
- 6-phospho-beta-D-glucosyl-(1,4)-D-glucose + H(2)O = D-glucose + D-glucose 6-phosphate
- Gene Name:
- bglA
- Uniprot ID:
- Q46829
- Molecular weight:
- 55361
Reactions
| 6-phospho-beta-D-glucosyl-(1,4)-D-glucose + H(2)O = D-glucose + D-glucose 6-phosphate. |
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for galactoside. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- mglC
- Uniprot ID:
- P23200
- Molecular weight:
- 35550
- General function:
- Involved in phosphoenolpyruvate-dependent sugar phosphotransferase system
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitant with their translocation across the cell membrane. This system is involved in mannose transport
- Gene Name:
- manY
- Uniprot ID:
- P69801
- Molecular weight:
- 27636
- General function:
- Involved in phosphoenolpyruvate-dependent sugar phosphotransferase system
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitant with their translocation across the cell membrane. This system is involved in mannose transport
- Gene Name:
- manZ
- Uniprot ID:
- P69805
- Molecular weight:
- 31303
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- This protein is involved in the active transport of galactose and glucose. It plays a role in the chemotaxis towards the two sugars by interacting with the trg chemoreceptor
- Gene Name:
- mglB
- Uniprot ID:
- P0AEE5
- Molecular weight:
- 35712
- General function:
- Involved in oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor
- Specific function:
- Aldose sugar dehydrogenase with broad substrate specificity. The physiological substrate is unknown. Can oxidize glucose to gluconolactone. Can also utilize D-arabinose, L- arabinose and 2-deoxy-glucose. Has higher activity towards oligomeric sugars, such as maltose, maltotriose or cellobiose. It may function to input sugar-derived electrons into the respiratory network
- Gene Name:
- yliI
- Uniprot ID:
- P75804
- Molecular weight:
- 41054
- General function:
- Involved in sugar:hydrogen symporter activity
- Specific function:
- General (non sugar-specific) component of the phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS). This major carbohydrate active-transport system catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. The phosphoryl group from phosphoenolpyruvate (PEP) is transferred to the phosphoryl carrier protein HPr by enzyme I. Phospho-HPr then transfers it to the permease (enzymes II/III)
- Gene Name:
- ptsH
- Uniprot ID:
- P0AA04
- Molecular weight:
- 9119
Reactions
| Protein HPr N(pi)-phospho-L-histidine + protein EIIA = protein HPr + protein EIIA N(tau)-phospho-L-histidine. |
- General function:
- carbohydrate metabolic process
- Specific function:
- Not Available
- Gene Name:
- lacZ
- Uniprot ID:
- G0ZKW2
- Molecular weight:
- 116482
Reactions
| = |
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex MglABC involved in galactose/methyl galactoside import. Responsible for energy coupling to the transport system (Probable)
- Gene Name:
- mglA
- Uniprot ID:
- P0AAG8
- Molecular weight:
- 56415
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in protein-N(PI)-phosphohistidine-sugar phosphotransferase activity
- Specific function:
- MalX encodes a phosphotransferase system enzyme II that can recognize glucose and maltose as substrates even though these sugars may not represent the natural substrates of the system
- Gene Name:
- malX
- Uniprot ID:
- P19642
- Molecular weight:
- 56627
Reactions
| Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate. |
- General function:
- Involved in protein-N(PI)-phosphohistidine-sugar phosphotransferase activity
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. This system is involved in glucose transport. This enzyme is also a chemoreceptor monitoring the environment for changes in sugar concentration
- Gene Name:
- ptsG
- Uniprot ID:
- P69786
- Molecular weight:
- 50676
Reactions
| Protein EIIB N(pi)-phospho-L-histidine/cysteine + sugar = protein EIIB + sugar phosphate. |
- General function:
- Involved in transmembrane transporter activity
- Specific function:
- Uptake of galactose across the boundary membrane with the concomitant transport of protons into the cell (symport system)
- Gene Name:
- galP
- Uniprot ID:
- P0AEP1
- Molecular weight:
- 50982
- General function:
- Involved in transmembrane transport
- Specific function:
- Involved in the efflux of sugars. The physiological role may be the detoxification of non-metabolizable sugar analogs. Can transport IPTG, lactose and glucose. Has broad substrate specificity, with preferences for glucosides or galactosides with alkyl or aryl substituents
- Gene Name:
- setA
- Uniprot ID:
- P31675
- Molecular weight:
- 42713
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for galactoside. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- mglC
- Uniprot ID:
- P23200
- Molecular weight:
- 35550
- General function:
- Involved in phosphoenolpyruvate-dependent sugar phosphotransferase system
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitant with their translocation across the cell membrane. This system is involved in mannose transport
- Gene Name:
- manY
- Uniprot ID:
- P69801
- Molecular weight:
- 27636
- General function:
- Involved in phosphoenolpyruvate-dependent sugar phosphotransferase system
- Specific function:
- The phosphoenolpyruvate-dependent sugar phosphotransferase system (PTS), a major carbohydrate active -transport system, catalyzes the phosphorylation of incoming sugar substrates concomitant with their translocation across the cell membrane. This system is involved in mannose transport
- Gene Name:
- manZ
- Uniprot ID:
- P69805
- Molecular weight:
- 31303
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Involved in porin activity
- Specific function:
- Involved in the transport of maltose and maltodextrins, indispensable for translocation of dextrins containing more than three glucosyl moieties. A hydrophobic path ("greasy slide") of aromatic residues serves to guide and select the sugars for transport through the channel. Also acts as a receptor for several bacteriophages including lambda
- Gene Name:
- lamB
- Uniprot ID:
- P02943
- Molecular weight:
- 49912
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- This protein is involved in the active transport of galactose and glucose. It plays a role in the chemotaxis towards the two sugars by interacting with the trg chemoreceptor
- Gene Name:
- mglB
- Uniprot ID:
- P0AEE5
- Molecular weight:
- 35712
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368