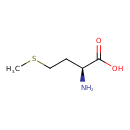
L-Methionine (ECMDB00696) (M2MDB000175)
Enzymes
- General function:
- Involved in nucleotide binding
- Specific function:
- Is required not only for elongation of protein synthesis but also for the initiation of all mRNA translation through initiator tRNA(fMet) aminoacylation
- Gene Name:
- metG
- Uniprot ID:
- P00959
- Molecular weight:
- 76254
Reactions
| ATP + L-methionine + tRNA(Met) = AMP + diphosphate + L-methionyl-tRNA(Met). |
- General function:
- Involved in transferase activity
- Specific function:
- An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate
- Gene Name:
- tyrB
- Uniprot ID:
- P04693
- Molecular weight:
- 43537
Reactions
| An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate. |
- General function:
- Involved in oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor
- Specific function:
- Could have an important function as a repair enzyme for proteins that have been inactivated by oxidation. Catalyzes the reversible oxidation-reduction of methionine sulfoxide in proteins to methionine
- Gene Name:
- msrA
- Uniprot ID:
- P0A744
- Molecular weight:
- 23315
Reactions
| Peptide-L-methionine + thioredoxin disulfide + H(2)O = peptide-L-methionine (S)-S-oxide + thioredoxin. |
| L-methionine + thioredoxin disulfide + H(2)O = L-methionine (S)-S-oxide + thioredoxin. |
- General function:
- Involved in peptide-methionine-(S)-S-oxide reductase activity
- Specific function:
- Peptide-L-methionine + thioredoxin disulfide + H(2)O = peptide-L-methionine (R)-S-oxide + thioredoxin
- Gene Name:
- msrB
- Uniprot ID:
- P0A746
- Molecular weight:
- 15451
Reactions
| Peptide-L-methionine + thioredoxin disulfide + H(2)O = peptide-L-methionine (R)-S-oxide + thioredoxin. |
- General function:
- Involved in methionine adenosyltransferase activity
- Specific function:
- Catalyzes the formation of S-adenosylmethionine from methionine and ATP. The overall synthetic reaction is composed of two sequential steps, AdoMet formation and the subsequent tripolyphosphate hydrolysis which occurs prior to release of AdoMet from the enzyme. Is essential for growth
- Gene Name:
- metK
- Uniprot ID:
- P0A817
- Molecular weight:
- 41951
Reactions
| ATP + L-methionine + H(2)O = phosphate + diphosphate + S-adenosyl-L-methionine. |
- General function:
- Involved in catalytic activity
- Specific function:
- Activation of pyruvate formate-lyase 1 under anaerobic conditions by generation of an organic free radical, using S- adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine
- Gene Name:
- pflA
- Uniprot ID:
- P0A9N4
- Molecular weight:
- 28204
Reactions
| S-adenosyl-L-methionine + dihydroflavodoxin + [formate C-acetyltransferase]-glycine = 5'-deoxyadenosine + L-methionine + flavodoxin semiquinone + [formate C-acetyltransferase]-glycin-2-yl radical. |
- General function:
- Involved in [formate-C-acetyltransferase]-activating enzyme activity
- Specific function:
- Activation of anaerobic ribonucleoside-triphosphate reductase under anaerobic conditions by generation of an organic free radical, using S-adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine
- Gene Name:
- nrdG
- Uniprot ID:
- P0A9N8
- Molecular weight:
- 17446
- General function:
- Involved in electron carrier activity
- Specific function:
- Efficient electron donor for the essential enzyme ribonucleotide reductase. Is also able to reduce the interchain disulfide bridges of insulin
- Gene Name:
- trxC
- Uniprot ID:
- P0AGG4
- Molecular weight:
- 15555
Reactions
| Protein dithiol + NAD(P)(+) = protein disulfide + NAD(P)H. |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the conversion of dethiobiotin (DTB) to biotin by the insertion of a sulfur atom into dethiobiotin via a radical- based mechanism
- Gene Name:
- bioB
- Uniprot ID:
- P12996
- Molecular weight:
- 38648
Reactions
| Dethiobiotin + sulfur + 2 S-adenosyl-L-methionine = biotin + 2 L-methionine + 2 5'-deoxyadenosine. |
- General function:
- Involved in methionine synthase activity
- Specific function:
- Catalyzes the transfer of a methyl group from methyl- cobalamin to homocysteine, yielding enzyme-bound cob(I)alamin and methionine. Subsequently, remethylates the cofactor using methyltetrahydrofolate
- Gene Name:
- metH
- Uniprot ID:
- P13009
- Molecular weight:
- 135996
Reactions
| 5-methyltetrahydrofolate + L-homocysteine = tetrahydrofolate + L-methionine. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- This enzyme may serve as a scavenger, allowing the cell to utilize biotin sulfoxide as a biotin source. It reduces a spontaneous oxidation product of biotin, D-biotin D-sulfoxide (BSO or BDS), back to biotin
- Gene Name:
- bisC
- Uniprot ID:
- P20099
- Molecular weight:
- 85850
- General function:
- Involved in 5-methyltetrahydropteroyltriglutamate-homocysteine S-methyltransferase activity
- Specific function:
- Catalyzes the transfer of a methyl group from 5- methyltetrahydrofolate to homocysteine resulting in methionine formation
- Gene Name:
- metE
- Uniprot ID:
- P25665
- Molecular weight:
- 84673
Reactions
| 5-methyltetrahydropteroyltri-L-glutamate + L-homocysteine = tetrahydropteroyltri-L-glutamate + L-methionine. |
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for D-methionine and the toxic methionine analog alpha-methyl- methionine. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- metI
- Uniprot ID:
- P31547
- Molecular weight:
- 23256
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes both the ATP-dependent activation of exogenously supplied lipoate to lipoyl-AMP and the transfer of the activated lipoyl onto the lipoyl domains of lipoate-dependent enzymes. Is also able to catalyze very poorly the transfer of lipoyl and octanoyl moiety from their acyl carrier protein
- Gene Name:
- lplA
- Uniprot ID:
- P32099
- Molecular weight:
- 37926
Reactions
| ATP + lipoate = diphosphate + lipoyl-AMP. |
| Lipoyl-AMP + protein = protein N(6)-(lipoyl)lysine + AMP. |
- General function:
- Involved in coproporphyrinogen oxidase activity
- Specific function:
- Anaerobic transformation of coproporphyrinogen-III into protoporphyrinogen-IX
- Gene Name:
- hemN
- Uniprot ID:
- P32131
- Molecular weight:
- 52729
Reactions
| Coproporphyrinogen-III + 2 S-adenosyl-L-methionine = protoporphyrinogen-IX + 2 CO(2) + 2 L-methionine + 2 5'-deoxyadenosine. |
- General function:
- Involved in iron-sulfur cluster binding
- Specific function:
- Activation of pyruvate formate-lyase 2 under anaerobic conditions by generation of an organic free radical, using S- adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine
- Gene Name:
- pflC
- Uniprot ID:
- P32675
- Molecular weight:
- 32429
Reactions
| S-adenosyl-L-methionine + dihydroflavodoxin + [formate C-acetyltransferase]-glycine = 5'-deoxyadenosine + L-methionine + flavodoxin semiquinone + [formate C-acetyltransferase]-glycin-2-yl radical. |
- General function:
- Involved in coproporphyrinogen oxidase activity
- Specific function:
- Not Available
- Gene Name:
- yggW
- Uniprot ID:
- P52062
- Molecular weight:
- 42584
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the radical-mediated insertion of two sulfur atoms into the C-6 and C-8 positions of the octanoyl moiety bound to the lipoyl domains of lipoate-dependent enzymes, thereby converting the octanoylated domains into lipoylated derivatives. Free octanoate is not a substrate for lipA
- Gene Name:
- lipA
- Uniprot ID:
- P60716
- Molecular weight:
- 36071
Reactions
| Protein N(6)-(octanoyl)lysine + 2 sulfur + 2 S-adenosyl-L-methionine = protein N(6)-(lipoyl)lysine + 2 L-methionine + 2 5'-deoxyadenosine. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Activation of pyruvate formate-lyase 2 under anaerobic conditions by generation of an organic free radical, using S- adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine
- Gene Name:
- ybiY
- Uniprot ID:
- P75794
- Molecular weight:
- 33038
Reactions
| S-adenosyl-L-methionine + dihydroflavodoxin + [formate C-acetyltransferase]-glycine = 5'-deoxyadenosine + L-methionine + flavodoxin semiquinone + [formate C-acetyltransferase]-glycin-2-yl radical. |
- General function:
- Involved in homocysteine S-methyltransferase activity
- Specific function:
- Catalyzes methyl transfer from S-methylmethionine or S- adenosylmethionine (less efficient) to homocysteine, selenohomocysteine and less efficiently selenocysteine
- Gene Name:
- mmuM
- Uniprot ID:
- Q47690
- Molecular weight:
- 33422
Reactions
| S-methyl-L-methionine + L-homocysteine = 2 L-methionine. |
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex MetNIQ involved in methionine import. Responsible for energy coupling to the transport system (Probable). It has also been shown to be involved in formyl-L-methionine transport
- Gene Name:
- metN
- Uniprot ID:
- P30750
- Molecular weight:
- 37788
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the rearrangement of 1-deoxy-D-xylulose 5- phosphate (DXP) to produce the thiazole phosphate moiety of thiamine. Sulfur is provided by the thiocarboxylate moiety of the carrier protein ThiS. In vitro, sulfur can be provided by H(2)S
- Gene Name:
- thiG
- Uniprot ID:
- P30139
- Molecular weight:
- 26896
Reactions
| 1-deoxy-D-xylulose 5-phosphate + 2-iminoacetate + thiocarboxy-adenylate-[sulfur-carrier protein ThiS] = 2-((2R,5Z)-2-carboxy-4-methylthiazol-5(2H)-ylidene)ethyl phosphate + [sulfur-carrier protein ThiS] + 2 H(2)O. |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the radical-mediated cleavage of tyrosine to dehydroglycine and p-cresol
- Gene Name:
- thiH
- Uniprot ID:
- P30140
- Molecular weight:
- 43320
Reactions
| L-tyrosine + S-adenosyl-L-methionine + reduced acceptor = 2-iminoacetate + 4-methylphenol + 5'-deoxyadenosine + L-methionine + acceptor + 2 H(+). |
- General function:
- Inorganic ion transport and metabolism
- Specific function:
- This protein is a component of a D-methionine permease, a binding protein-dependent, ATP-driven transport system
- Gene Name:
- metQ
- Uniprot ID:
- P28635
- Molecular weight:
- 29432
- General function:
- Involved in thiamine biosynthetic process
- Specific function:
- Catalyzes the synthesis of the hydroxymethylpyrimidine phosphate (HMP-P) moiety of thiamine from aminoimidazole ribotide (AIR) in a radical S-adenosyl-L-methionine (SAM)-dependent reaction
- Gene Name:
- thiC
- Uniprot ID:
- P30136
- Molecular weight:
- 70850
Reactions
| 5-amino-1-(5-phospho-D-ribosyl)imidazole + S-adenosyl-L-methionine = 4-amino-2-methyl-5-phosphomethylpyrimidine + 5'-deoxyadenosine + L-methionine + formate + CO. |
- General function:
- Involved in electron carrier activity
- Specific function:
- Participates in various redox reactions through the reversible oxidation of its active center dithiol to a disulfide and catalyzes dithiol-disulfide exchange reactions
- Gene Name:
- trxA
- Uniprot ID:
- P0AA25
- Molecular weight:
- 11807
- General function:
- Not Available
- Specific function:
- Not Available
- Gene Name:
- yncA
- Uniprot ID:
- P76112
- Molecular weight:
- Not Available
- General function:
- Translation, ribosomal structure and biogenesis
- Specific function:
- Transfers and isomerizes the ribose moiety from AdoMet to the 7-aminomethyl group of 7-deazaguanine (preQ1-tRNA) to give epoxyqueuosine (oQ-tRNA)
- Gene Name:
- queA
- Uniprot ID:
- P0A7F9
- Molecular weight:
- 39430
Reactions
| S-adenosylmethionine + 7-aminomethyl-7-deazaguanosine = methionine + adenine + epoxyqueuosine. |
- General function:
- Involved in 4 iron, 4 sulfur cluster binding
- Specific function:
- Specifically methylates position 2 of adenine 2503 in 23S rRNA
- Gene Name:
- rlmN
- Uniprot ID:
- P36979
- Molecular weight:
- 43085
Reactions
| 2 S-adenosyl-L-methionine + adenine(2503) in 23S rRNA = S-adenosyl-L-homocysteine + L-methionine + 5'-deoxyadenosine + 2-methyladenine(2503) in 23S rRNA. |
- General function:
- tRNA methylthiolation
- Specific function:
- Catalyzes the methylthiolation of N6-(dimethylallyl)adenosine (i(6)A), leading to the formation of 2-methylthio-N6-(dimethylallyl)adenosine (ms(2)i(6)A) at position 37 in tRNAs that read codons beginning with uridine.
- Gene Name:
- miaB
- Uniprot ID:
- P0AEI1
- Molecular weight:
- 53662
Reactions
| N(6)-dimethylallyladenine(37) in tRNA + sulfur-(sulfur carrier) + 2 S-adenosyl-L-methionine = 2-methylthio-N(6)-dimethylallyladenine(37) in tRNA + S-adenosyl-L-homocysteine + (sulfur carrier) + L-methionine + 5'-deoxyadenosine |
- General function:
- RNA modification
- Specific function:
- Catalyzes the methylthiolation of the residue Asp-89 of ribosomal protein S12.
- Gene Name:
- rimO
- Uniprot ID:
- P0AEI4
- Molecular weight:
- 49581
Reactions
| L-aspartate-[ribosomal protein S12] + sulfur-(sulfur carrier) + 2 S-adenosyl-L-methionine = 3-methylthio-L-aspartate-[ribosomal protein S12] + S-adenosyl-L-homocysteine + (sulfur carrier) + L-methionine + 5'-deoxyadenosine |
Transporters
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for D-methionine and the toxic methionine analog alpha-methyl- methionine. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- metI
- Uniprot ID:
- P31547
- Molecular weight:
- 23256
- General function:
- Involved in nucleotide binding
- Specific function:
- Probably part of a binding-protein-dependent transport system yecCS for an amino acid. Probably responsible for energy coupling to the transport system
- Gene Name:
- yecC
- Uniprot ID:
- P37774
- Molecular weight:
- 27677
- General function:
- Involved in transporter activity
- Specific function:
- Probably part of the binding-protein-dependent transport system yecCS for an amino acid; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- yecS
- Uniprot ID:
- P0AFT2
- Molecular weight:
- 24801
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex MetNIQ involved in methionine import. Responsible for energy coupling to the transport system (Probable). It has also been shown to be involved in formyl-L-methionine transport
- Gene Name:
- metN
- Uniprot ID:
- P30750
- Molecular weight:
- 37788
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Inorganic ion transport and metabolism
- Specific function:
- This protein is a component of a D-methionine permease, a binding protein-dependent, ATP-driven transport system
- Gene Name:
- metQ
- Uniprot ID:
- P28635
- Molecular weight:
- 29432
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368