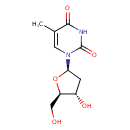
Thymidine (ECMDB00273) (M2MDB000114)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 10:25:22 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-09-13 12:56:07 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Thymidine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Thymidine is non-toxic and is a naturally occurring compound that exists in all living organisms and DNA viruses. 25% of DNA is composed of thymidine. RNA does not have thymidine and has uridine instead. Thymidine is a chemical compound which is a pyrimidine nucleoside. Thymidine is the DNA base T, which pairs with adenosine in double stranded DNA. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C10H14N2O5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 242.2286 Monoisotopic: 242.090271568 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | IQFYYKKMVGJFEH-XLPZGREQSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C10H14N2O5/c1-5-3-12(10(16)11-9(5)15)8-2-6(14)7(4-13)17-8/h3,6-8,13-14H,2,4H2,1H3,(H,11,15,16)/t6-,7+,8+/m0/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 50-89-5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 1-[(2R,4S,5R)-4-hydroxy-5-(hydroxymethyl)oxolan-2-yl]-5-methyl-1,2,3,4-tetrahydropyrimidine-2,4-dione | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | thymidine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | CC1=CN([C@H]2C[C@H](O)[C@@H](CO)O2)C(=O)NC1=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as pyrimidine 2'-deoxyribonucleosides. Pyrimidine 2'-deoxyribonucleosides are compounds consisting of a pyrimidine linked to a ribose which lacks a hydroxyl group at position 2. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Nucleosides, nucleotides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pyrimidine nucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Pyrimidine 2'-deoxyribonucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Pyrimidine 2'-deoxyribonucleosides | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | 5-Thymidylic acid + Water > Phosphate + Thymidine Adenosine triphosphate + Thymidine <> ADP + 5-Thymidylic acid + Hydrogen ion Phosphate + Thymidine <> Deoxyribose 1-phosphate + Thymine Adenosine triphosphate + Thymidine <> ADP + 5-Thymidylic acid 5-Thymidylic acid + Water <> Thymidine + Phosphate Thymidine + Adenosine triphosphate > Hydrogen ion + 5-Thymidylic acid + ADP Thymidine + Phosphate <> deoxyribose-1-phosphate + Thymine Adenosine triphosphate + Thymidine > ADP + 5-Thymidylic acid Thymidine + Inorganic phosphate > Thymine + 2-deoxy-alpha-D-ribose 1-phosphate Deoxyribose 1-phosphate + Thymine > Phosphate + Thymidine 5 5-Thymidylic acid + Water > Phosphate + Thymidine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Herdewijn, P.; Kerremans, L.; Wigerinck, P.; Vandendriessche, F.; Van Aerschot, A. Synthesis of thymidine from 5-iodo-2'-deoxyuridine. Tetrahedron Letters (1991), 32(34), 4397-400. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in hydrolase activity
- Specific function:
- Degradation of external UDP-glucose to uridine monophosphate and glucose-1-phosphate, which can then be used by the cell
- Gene Name:
- ushA
- Uniprot ID:
- P07024
- Molecular weight:
- 60824
Reactions
| UDP-sugar + H(2)O = UMP + alpha-D-aldose 1-phosphate. |
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in transferase activity, transferring glycosyl groups
- Specific function:
- The enzymes which catalyze the reversible phosphorolysis of pyrimidine nucleosides are involved in the degradation of these compounds and in their utilization as carbon and energy sources, or in the rescue of pyrimidine bases for nucleotide synthesis
- Gene Name:
- deoA
- Uniprot ID:
- P07650
- Molecular weight:
- 47207
Reactions
| Thymidine + phosphate = thymine + 2-deoxy-alpha-D-ribose 1-phosphate. |
- General function:
- Involved in hydrolase activity
- Specific function:
- Nucleotidase with a broad substrate specificity as it can dephosphorylate various ribo- and deoxyribonucleoside 5'- monophosphates and ribonucleoside 3'-monophosphates with highest affinity to 3'-AMP. Also hydrolyzes polyphosphate (exopolyphosphatase activity) with the preference for short-chain- length substrates (P20-25). Might be involved in the regulation of dNTP and NTP pools, and in the turnover of 3'-mononucleotides produced by numerous intracellular RNases (T1, T2, and F) during the degradation of various RNAs. Also plays a significant physiological role in stress-response and is required for the survival of E.coli in stationary growth phase
- Gene Name:
- surE
- Uniprot ID:
- P0A840
- Molecular weight:
- 26900
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| A 3'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
| (Polyphosphate)(n) + H(2)O = (polyphosphate)(n-1) + phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows high phosphatase activity toward three nucleoside 5'-monophosphates, UMP, dUMP, and dTMP, and very low activity against TDP, IMP, UDP, GMP, dGMP, AMP, dAMP, and 6- phosphogluconate. Is strictly specific to substrates with 5'- phosphates and shows no activity against nucleoside 2'- or 3'- monophosphates. Might be involved in the pyrimidine nucleotide substrate cycles
- Gene Name:
- yjjG
- Uniprot ID:
- P0A8Y1
- Molecular weight:
- 25300
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Uniprot ID:
- P0AE22
- Molecular weight:
- 26103
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in thymidine kinase activity
- Specific function:
- Phosphorylates both thymidine and deoxyuridine
- Gene Name:
- tdk
- Uniprot ID:
- P23331
- Molecular weight:
- 23456
Reactions
| ATP + thymidine = ADP + thymidine 5'-phosphate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Nucleotidase that shows strict specificity toward deoxyribonucleoside 5'-monophosphates and does not dephosphorylate 5'-ribonucleotides or ribonucleoside 3'-monophosphates. Might be involved in the regulation of all dNTP pools in E.coli
- Gene Name:
- yfbR
- Uniprot ID:
- P76491
- Molecular weight:
- 22708
Reactions
| A 5'-ribonucleotide + H(2)O = a ribonucleoside + phosphate. |
Transporters
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Nucleoside transport
- Gene Name:
- xapB
- Uniprot ID:
- P45562
- Molecular weight:
- 46139
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Transports nucleosides with a high affinity except guanosine and deoxyguanosine. Driven by a proton motive force
- Gene Name:
- nupC
- Uniprot ID:
- P0AFF2
- Molecular weight:
- 43475
- General function:
- Involved in nucleoside transmembrane transporter activity
- Specific function:
- Transports nucleosides with a high affinity. Driven by a proton motive force
- Gene Name:
- nupG
- Uniprot ID:
- P0AFF4
- Molecular weight:
- 46389
- General function:
- Involved in nucleoside:sodium symporter activity
- Specific function:
- Nucleoside transporter
- Gene Name:
- nupX
- Uniprot ID:
- P33021
- Molecular weight:
- 43409
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368