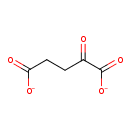
alpha-Ketoglutarate (ECMDB02812) (M2MDB000466)
Enzymes
- General function:
- Involved in oxidoreductase activity
- Specific function:
- L-glutamate + H(2)O + NADP(+) = 2-oxoglutarate + NH(3) + NADPH
- Gene Name:
- gdhA
- Uniprot ID:
- P00370
- Molecular weight:
- 48581
Reactions
| L-glutamate + H(2)O + NADP(+) = 2-oxoglutarate + NH(3) + NADPH. |
- General function:
- Involved in transferase activity
- Specific function:
- L-aspartate + 2-oxoglutarate = oxaloacetate + L-glutamate
- Gene Name:
- aspC
- Uniprot ID:
- P00509
- Molecular weight:
- 43573
Reactions
| L-aspartate + 2-oxoglutarate = oxaloacetate + L-glutamate. |
- General function:
- Involved in transferase activity
- Specific function:
- An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate
- Gene Name:
- tyrB
- Uniprot ID:
- P04693
- Molecular weight:
- 43537
Reactions
| An aromatic amino acid + 2-oxoglutarate = an aromatic oxo acid + L-glutamate. |
- General function:
- Involved in transferase activity
- Specific function:
- L-histidinol phosphate + 2-oxoglutarate = 3- (imidazol-4-yl)-2-oxopropyl phosphate + L-glutamate
- Gene Name:
- hisC
- Uniprot ID:
- P06986
- Molecular weight:
- 39360
Reactions
| L-histidinol phosphate + 2-oxoglutarate = 3-(imidazol-4-yl)-2-oxopropyl phosphate + L-glutamate. |
- General function:
- Involved in magnesium ion binding
- Specific function:
- Isocitrate + NADP(+) = 2-oxoglutarate + CO(2) + NADPH
- Gene Name:
- icd
- Uniprot ID:
- P08200
- Molecular weight:
- 45756
Reactions
| Isocitrate + NADP(+) = 2-oxoglutarate + CO(2) + NADPH. |
- General function:
- Involved in catalytic activity
- Specific function:
- 2 L-glutamate + NADP(+) = L-glutamine + 2- oxoglutarate + NADPH
- Gene Name:
- gltB
- Uniprot ID:
- P09831
- Molecular weight:
- 166708
Reactions
| 2 L-glutamate + NADP(+) = L-glutamine + 2-oxoglutarate + NADPH. |
- General function:
- Involved in iron-sulfur cluster binding
- Specific function:
- 2 L-glutamate + NADP(+) = L-glutamine + 2- oxoglutarate + NADPH
- Gene Name:
- gltD
- Uniprot ID:
- P09832
- Molecular weight:
- 52015
Reactions
| 2 L-glutamate + NADP(+) = L-glutamine + 2-oxoglutarate + NADPH. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Lipoamide dehydrogenase is a component of the glycine cleavage system as well as of the alpha-ketoacid dehydrogenase complexes
- Gene Name:
- lpdA
- Uniprot ID:
- P0A9P0
- Molecular weight:
- 50688
Reactions
| Protein N(6)-(dihydrolipoyl)lysine + NAD(+) = protein N(6)-(lipoyl)lysine + NADH. |
- General function:
- Involved in oxoglutarate dehydrogenase (succinyl-transferring) activity
- Specific function:
- The 2-oxoglutarate dehydrogenase complex catalyzes the overall conversion of 2-oxoglutarate to succinyl-CoA and CO(2). It contains multiple copies of three enzymatic components:2- oxoglutarate dehydrogenase (E1), dihydrolipoamide succinyltransferase (E2) and lipoamide dehydrogenase (E3)
- Gene Name:
- sucA
- Uniprot ID:
- P0AFG3
- Molecular weight:
- 105061
Reactions
| 2-oxoglutarate + [dihydrolipoyllysine-residue succinyltransferase] lipoyllysine = [dihydrolipoyllysine-residue succinyltransferase] S-succinyldihydrolipoyllysine + CO(2). |
10. Dihydrolipoyllysine-residue succinyltransferase component of 2-oxoglutarate dehydrogenase complex
- General function:
- Involved in transferase activity, transferring acyl groups
- Specific function:
- The 2-oxoglutarate dehydrogenase complex catalyzes the overall conversion of 2-oxoglutarate to succinyl-CoA and CO(2). It contains multiple copies of three enzymatic components:2- oxoglutarate dehydrogenase (E1), dihydrolipoamide succinyltransferase (E2) and lipoamide dehydrogenase (E3)
- Gene Name:
- sucB
- Uniprot ID:
- P0AFG6
- Molecular weight:
- 44011
Reactions
| Succinyl-CoA + enzyme N(6)-(dihydrolipoyl)lysine = CoA + enzyme N(6)-(S-succinyldihydrolipoyl)lysine. |
- General function:
- Involved in thiamine pyrophosphate binding
- Specific function:
- Catalyzes the thiamine diphosphate-dependent decarboxylation of 2-oxoglutarate and the subsequent addition of the resulting succinic semialdehyde-thiamine pyrophosphate anion to isochorismate to yield 2-succinyl-5-enolpyruvyl-6-hydroxy-3- cyclohexene-1-carboxylate (SEPHCHC)
- Gene Name:
- menD
- Uniprot ID:
- P17109
- Molecular weight:
- 61367
Reactions
| Isochorismate + 2-oxoglutarate = 5-enolpyruvoyl-6-hydroxy-2-succinyl-cyclohex-3-ene-1-carboxylate + CO(2). |
- General function:
- Involved in transaminase activity
- Specific function:
- Involved in both the arginine and lysine biosynthetic pathways
- Gene Name:
- argD
- Uniprot ID:
- P18335
- Molecular weight:
- 43767
Reactions
| N(2)-acetyl-L-ornithine + 2-oxoglutarate = N-acetyl-L-glutamate 5-semialdehyde + L-glutamate. |
| N-succinyl-L-2,6-diaminoheptanedioate + 2-oxoglutarate = N-succinyl-2-L-amino-6-oxoheptanedioate + L-glutamate. |
- General function:
- Involved in 4-aminobutyrate transaminase activity
- Specific function:
- 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate
- Gene Name:
- gabT
- Uniprot ID:
- P22256
- Molecular weight:
- 45774
Reactions
| 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate. |
| (S)-3-amino-2-methylpropanoate + 2-oxoglutarate = 2-methyl-3-oxopropanoate + L-glutamate. |
- General function:
- Involved in metabolic process
- Specific function:
- Catalyzes the reversible conversion of 3- phosphohydroxypyruvate to phosphoserine and of 3-hydroxy-2-oxo-4- phosphonooxybutanoate to phosphohydroxythreonine. Is involved in both pyridoxine and serine biosynthesis
- Gene Name:
- serC
- Uniprot ID:
- P23721
- Molecular weight:
- 39783
Reactions
| O-phospho-L-serine + 2-oxoglutarate = 3-phosphonooxypyruvate + L-glutamate. |
| 4-phosphonooxy-L-threonine + 2-oxoglutarate = (3R)-3-hydroxy-2-oxo-4-phosphonooxybutanoate + L-glutamate. |
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Catalyzes the conversion of taurine and alpha ketoglutarate to sulfite, aminoacetaldehyde and succinate. Required for the utilization of taurine (2-aminoethanesulfonic acid) as an alternative sulfur source. Pentane-sulfonic acid, 3- (N-morpholino)propanesulfonic acid and 1,3-dioxo-2- isoindolineethanesulfonic acid are also substrates for this enzyme
- Gene Name:
- tauD
- Uniprot ID:
- P37610
- Molecular weight:
- 32409
Reactions
| Taurine + 2-oxoglutarate + O(2) = sulfite + aminoacetaldehyde + succinate + CO(2). |
- General function:
- Involved in transaminase activity
- Specific function:
- Catalyzes the aminotransferase reaction from putrescine to 2-oxoglutarate, leading to glutamate and 4-aminobutanal, which spontaneously cyclizes to form 1-pyrroline. Is also able to transaminate cadaverine and, in lower extent, spermidine, but not ornithine. Alpha-ketobutyrate and pyruvate can also act as amino acceptors, although much less efficiently
- Gene Name:
- patA
- Uniprot ID:
- P42588
- Molecular weight:
- 49661
Reactions
| Putrescine + 2-oxoglutarate = L-glutamate + 1-pyrroline + H(2)O. |
- General function:
- Involved in 4-aminobutyrate transaminase activity
- Specific function:
- Involved in the breakdown of putrescine via transamination of gamma-aminobutyrate
- Gene Name:
- puuE
- Uniprot ID:
- P50457
- Molecular weight:
- 44729
Reactions
| 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate. |
- General function:
- Involved in transaminase activity
- Specific function:
- Catalyzes the transamination of N(2)-succinylornithine and alpha-ketoglutarate into N(2)-succinylglutamate semialdehyde and glutamate. Can also act as an acetylornithine aminotransferase
- Gene Name:
- astC
- Uniprot ID:
- P77581
- Molecular weight:
- 43665
Reactions
| N(2)-succinyl-L-ornithine + 2-oxoglutarate = N-succinyl-L-glutamate 5-semialdehyde + L-glutamate. |
- General function:
- Involved in transaminase activity
- Specific function:
- Catalyzes the conversion of UDP-4-keto-arabinose (UDP- Ara4O) to UDP-4-amino-4-deoxy-L-arabinose (UDP-L-Ara4N). The modified arabinose is attached to lipid A and is required for resistance to polymyxin and cationic antimicrobial peptides
- Gene Name:
- arnB
- Uniprot ID:
- P77690
- Molecular weight:
- 41649
Reactions
| UDP-4-amino-4-deoxy-beta-L-arabinose + 2-oxoglutarate = UDP-beta-L-threo-pentapyranos-4-ulose + glutamate. |
- General function:
- Involved in 1-aminocyclopropane-1-carboxylate synthase activity
- Specific function:
- Specific function unknown
- Gene Name:
- yfbQ
- Uniprot ID:
- P0A959
- Molecular weight:
- 45517
Reactions
| L-alanine + 2-oxoglutarate = pyruvate + L-glutamate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Acts on leucine, isoleucine and valine
- Gene Name:
- ilvE
- Uniprot ID:
- P0AB80
- Molecular weight:
- 34093
Reactions
| L-leucine + 2-oxoglutarate = 4-methyl-2-oxopentanoate + L-glutamate. |
| L-isoleucine + 2-oxoglutarate = (S)-3-methyl-2-oxopentanoate + L-glutamate. |
| L-valine + 2-oxoglutarate = 3-methyl-2-oxobutanoate + L-glutamate. |
- General function:
- Involved in catalytic activity
- Specific function:
- Involved in ECA elongation
- Gene Name:
- rffA
- Uniprot ID:
- P27833
- Molecular weight:
- 41901
- General function:
- Amino acid transport and metabolism
- Specific function:
- Specific function unknown
- Gene Name:
- yfdZ
- Uniprot ID:
- P77434
- Molecular weight:
- 46216
Reactions
| L-alanine + 2-oxoglutarate = pyruvate + L-glutamate. |
Transporters
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of alpha-ketoglutarate across the boundary membrane with the concomitant import of a cation (symport system)
- Gene Name:
- kgtP
- Uniprot ID:
- P0AEX3
- Molecular weight:
- 47052
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368