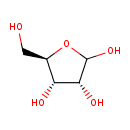
Ribose (ECMDB00283) (M2MDB000116)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 10:25:29 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-09-13 15:15:18 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | Ribose | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Ribose is a pentose that is actively used in many biological systems usually in its D-form. Ribose constitutes the backbone of RNA, a biopolymer that is the basis of genetic transcription. It is related to deoxyribose, as found in DNA. Once phosphorylated, ribose can become a subunit of ATP, NADH, and several other compounds that are critical to metabolism like the secondary messengers cAMP and cGMP. In cells D-ribose must be phosphorylated before it can be used. Ribokinase catalyzes this reaction by converting D-ribose to D-ribose 5-phosphate. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C5H10O5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 150.1299 Monoisotopic: 150.05282343 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | HMFHBZSHGGEWLO-SOOFDHNKSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C5H10O5/c6-1-2-3(7)4(8)5(9)10-2/h2-9H,1H2/t2-,3-,4-,5?/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 50-69-1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (3R,4S,5R)-5-(hydroxymethyl)oxolane-2,3,4-triol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | D-ribofuranoside | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | OC[C@H]1OC(O)[C@H](O)[C@@H]1O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as pentoses. These are monosaccharides in which the carbohydrate moiety contains five carbon atoms. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic oxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Carbohydrates and carbohydrate conjugates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Pentoses | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + Water + Ribose > ADP + Hydrogen ion + Phosphate + Ribose Adenosine triphosphate + Water + Ribose > ADP + Hydrogen ion + Phosphate + Ribose Cytidine + Water > Cytosine + Ribose Water + Uridine > Ribose + Uracil Adenosine + Water > Adenine + Ribose Water + Inosine > Hypoxanthine + Ribose Water + Xanthosine > Ribose + Xanthine Water + D-Ribose-5-phosphate > Phosphate + Ribose Adenosine triphosphate + Ribose <> ADP + Hydrogen ion + D-Ribose-5-phosphate Adenosine triphosphate + Ribose <> ADP + D-Ribose-5-phosphate Adenosine + Water <> Adenine + Ribose Guanosine + Water <> Guanine + Ribose Inosine + Water <> Hypoxanthine + Ribose Cytidine + Water <> Cytosine + Ribose Adenosine triphosphate + Ribose > ADP + D-Ribose-5-phosphate A pyrimidine nucleoside + Water > Ribose + a pyrimidine base Pyrimidine nucleoside + Water <> Ribose + Pyrimidine Ribose + Adenosine triphosphate > D-Ribose-5-phosphate + ADP + Hydrogen ion Adenosine triphosphate + Ribose <> ADP + Hydrogen ion + D-Ribose-5-phosphate Adenosine triphosphate + Ribose <> ADP + Hydrogen ion + D-Ribose-5-phosphate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Park, Yong-Cheol; Choi, Jin-Ho; Bennett, George N.; Seo, Jin-Ho. Characterization of D-ribose biosynthesis in Bacillus subtilis JY200 deficient in transketolase gene. Journal of Biotechnology (2006), 121(4), 508-516. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex RbsABCD involved in ribose import. Responsible for energy coupling to the transport system
- Gene Name:
- rbsA
- Uniprot ID:
- P04983
- Molecular weight:
- 55041
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in phosphotransferase activity, alcohol group as acceptor
- Specific function:
- ATP + D-ribose = ADP + D-ribose 5-phosphate
- Gene Name:
- rbsK
- Uniprot ID:
- P0A9J6
- Molecular weight:
- 32290
Reactions
| ATP + D-ribose = ADP + D-ribose 5-phosphate. |
- General function:
- Involved in acid phosphatase activity
- Specific function:
- Dephosphorylates several organic phosphomonoesters and catalyzes the transfer of low-energy phosphate groups from phosphomonoesters to hydroxyl groups of various organic compounds. Preferentially acts on aryl phosphoesters. Might function as a broad-spectrum dephosphorylating enzyme able to scavenge both 3'- and 5'-nucleotides and also additional organic phosphomonoesters
- Gene Name:
- aphA
- Uniprot ID:
- P0AE22
- Molecular weight:
- 26103
Reactions
| A phosphate monoester + H(2)O = an alcohol + phosphate. |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes both purine and pyrimidine ribonucleosides with a broad-substrate specificity with decreasing activity in the order uridine, xanthosine, inosine, adenosine, cytidine, guanosine
- Gene Name:
- rihC
- Uniprot ID:
- P22564
- Molecular weight:
- 32560
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex AlsBAC involved in D-allose import. Probably responsible for energy coupling to the transport system
- Gene Name:
- alsA
- Uniprot ID:
- P32721
- Molecular weight:
- 56744
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes cytidine or uridine to ribose and cytosine or uracil, respectively. Has a clear preference for cytidine over uridine. Strictly specific for ribonucleosides. Has a low but significant activity for the purine nucleoside xanthosine
- Gene Name:
- rihB
- Uniprot ID:
- P33022
- Molecular weight:
- 33748
Reactions
| A pyrimidine nucleoside + H(2)O = D-ribose + a pyrimidine base. |
- General function:
- Involved in catalytic activity
- Specific function:
- Catalyzes the hydrolysis of sugar phosphate to sugar and inorganic phosphate. Has a wide substrate specificity catalyzing the hydrolysis of fructose-1-P most efficiently, but it remains uncertain if this is the real substrate in vivo
- Gene Name:
- supH
- Uniprot ID:
- P75792
- Molecular weight:
- 30413
Reactions
| Sugar phosphate + H(2)O = sugar + phosphate. |
- General function:
- Involved in nucleotide binding
- Specific function:
- Specific function unknown
- Gene Name:
- ytfR
- Uniprot ID:
- Q6BEX0
- Molecular weight:
- 55259
- General function:
- Involved in transporter activity
- Specific function:
- Probably part of a binding-protein-dependent transport system. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- yjfF
- Uniprot ID:
- P37772
- Molecular weight:
- 34977
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Specific function unknown
- Gene Name:
- ytfQ
- Uniprot ID:
- P39325
- Molecular weight:
- 34344
- General function:
- Involved in transporter activity
- Specific function:
- Probably part of a binding-protein-dependent transport system. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- ytfT
- Uniprot ID:
- P39328
- Molecular weight:
- 35659
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for ribose. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- rbsC
- Uniprot ID:
- P0AGI1
- Molecular weight:
- 33452
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system AlsBAC for D-allose; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- alsC
- Uniprot ID:
- P32720
- Molecular weight:
- 34315
- General function:
- Involved in intramolecular lyase activity
- Specific function:
- Catalyzes the interconversion of beta-pyran and beta- furan forms of D-ribose. It also catalyzes the conversion between beta-allofuranose and beta-allopyranose
- Gene Name:
- rbsD
- Uniprot ID:
- P04982
- Molecular weight:
- 15292
Reactions
| Beta-D-ribopyranose = beta-D-ribofuranose. |
| Beta-D-allopyranose = beta-D-allofuranose. |
- General function:
- Involved in hydrolase activity, hydrolyzing N-glycosyl compounds
- Specific function:
- Hydrolyzes with equal efficiency cytidine or uridine to ribose and cytosine or uracil, respectively
- Gene Name:
- rihA
- Uniprot ID:
- P41409
- Molecular weight:
- 33823
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Part of the binding-protein-dependent transport system AlsBAC for D-allose
- Gene Name:
- alsB
- Uniprot ID:
- P39265
- Molecular weight:
- 32910
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Involved in the high-affinity D-ribose membrane transport system and also serves as the primary chemoreceptor for chemotaxis
- Gene Name:
- rbsB
- Uniprot ID:
- P02925
- Molecular weight:
- 30950
Transporters
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex RbsABCD involved in ribose import. Responsible for energy coupling to the transport system
- Gene Name:
- rbsA
- Uniprot ID:
- P04983
- Molecular weight:
- 55041
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in nucleotide binding
- Specific function:
- Part of the ABC transporter complex AlsBAC involved in D-allose import. Probably responsible for energy coupling to the transport system
- Gene Name:
- alsA
- Uniprot ID:
- P32721
- Molecular weight:
- 56744
Reactions
| ATP + H(2)O + monosaccharide(Out) = ADP + phosphate + monosaccharide(In). |
- General function:
- Involved in nucleotide binding
- Specific function:
- Specific function unknown
- Gene Name:
- ytfR
- Uniprot ID:
- Q6BEX0
- Molecular weight:
- 55259
- General function:
- Involved in transporter activity
- Specific function:
- Probably part of a binding-protein-dependent transport system. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- yjfF
- Uniprot ID:
- P37772
- Molecular weight:
- 34977
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Specific function unknown
- Gene Name:
- ytfQ
- Uniprot ID:
- P39325
- Molecular weight:
- 34344
- General function:
- Involved in transporter activity
- Specific function:
- Probably part of a binding-protein-dependent transport system. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- ytfT
- Uniprot ID:
- P39328
- Molecular weight:
- 35659
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system for ribose. Probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- rbsC
- Uniprot ID:
- P0AGI1
- Molecular weight:
- 33452
- General function:
- Involved in transporter activity
- Specific function:
- Part of the binding-protein-dependent transport system AlsBAC for D-allose; probably responsible for the translocation of the substrate across the membrane
- Gene Name:
- alsC
- Uniprot ID:
- P32720
- Molecular weight:
- 34315
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Part of the binding-protein-dependent transport system AlsBAC for D-allose
- Gene Name:
- alsB
- Uniprot ID:
- P39265
- Molecular weight:
- 32910
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Carbohydrate transport and metabolism
- Specific function:
- Involved in the high-affinity D-ribose membrane transport system and also serves as the primary chemoreceptor for chemotaxis
- Gene Name:
- rbsB
- Uniprot ID:
- P02925
- Molecular weight:
- 30950
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368