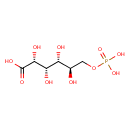
6-Phosphogluconic acid (ECMDB01316) (M2MDB000336)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 13:49:48 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-09-13 12:56:10 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | 6-Phosphogluconic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | 6-Phosphogluconic acid is an intermediate in the Pentose phosphate pathway (KEGG) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C6H13O10P | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 276.1352 Monoisotopic: 276.024633148 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | BIRSGZKFKXLSJQ-SQOUGZDYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C6H13O10P/c7-2(1-16-17(13,14)15)3(8)4(9)5(10)6(11)12/h2-5,7-10H,1H2,(H,11,12)(H2,13,14,15)/t2-,3-,4+,5-/m1/s1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 921-62-0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | (2R,3S,4R,5R)-2,3,4,5-tetrahydroxy-6-(phosphonooxy)hexanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | 6-phosphogluconic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | O[C@H](COP(O)(O)=O)[C@@H](O)[C@H](O)[C@@H](O)C(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as monosaccharide phosphates. These are monosaccharides comprising a phosphated group linked to the carbohydrate unit. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic oxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Organooxygen compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Carbohydrates and carbohydrate conjugates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Monosaccharide phosphates | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | -3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | Adenosine triphosphate + Gluconic acid <> 6-Phosphogluconic acid + ADP + Hydrogen ion 6-Phosphonoglucono-D-lactone + Water <> 6-Phosphogluconic acid + Hydrogen ion 6-Phosphogluconic acid <> 2-Keto-3-deoxy-6-phosphogluconic acid + Water 6-Phosphogluconic acid + NADP <> Carbon dioxide + NADPH + D-Ribulose 5-phosphate + Hydrogen ion 6-Phosphogluconic acid + NADP <> D-Ribulose 5-phosphate + Carbon dioxide + NADPH + Hydrogen ion Adenosine triphosphate + Gluconic acid <> ADP + 6-Phosphogluconic acid 6-Phosphonoglucono-D-lactone + Water <> 6-Phosphogluconic acid NAD(P)<sup>+</sup> + 6-Phosphogluconic acid > NAD(P)H + D-Ribulose 5-phosphate + Carbon dioxide 6-Phosphonoglucono-D-lactone + Water > Hydrogen ion + 6-Phosphogluconic acid Adenosine triphosphate + Gluconic acid > Hydrogen ion + ADP + 6-Phosphogluconic acid 6-Phosphogluconic acid > 2-Keto-3-deoxy-6-phosphogluconic acid + Water 6-Phosphogluconic acid + Water > Gluconic acid + Phosphate 6-Phosphogluconic acid + NADP > D-Ribulose 5-phosphate + Carbon dioxide + NADPH 6-Phosphonoglucono-D-lactone + Water > 6-Phosphogluconic acid Adenosine triphosphate + Gluconic acid > ADP + 6-Phosphogluconic acid 6-Phosphogluconic acid + NADP > D-Ribulose 5-phosphate + Carbon dioxide + NADPH + NADPH 6 6-Phosphonoglucono-D-lactone + Water <>6 6-Phosphogluconic acid + Hydrogen ion 6 6-Phosphonoglucono-D-lactone + Water <>6 6-Phosphogluconic acid 6 6-Phosphogluconic acid <>2 2-Keto-3-deoxy-6-phosphogluconic acid + Water 6 6-Phosphonoglucono-D-lactone + Water <>6 6-Phosphogluconic acid + Hydrogen ion | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Find out more about how we convert literature concentrations. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Download (PDF) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in oxidation-reduction process
- Specific function:
- Catalyzes the oxidative decarboxylation of 6- phosphogluconate to ribulose 5-phosphate and CO(2), with concomitant reduction of NADP to NADPH
- Gene Name:
- gnd
- Uniprot ID:
- P00350
- Molecular weight:
- 51481
Reactions
| 6-phospho-D-gluconate + NADP(+) = D-ribulose 5-phosphate + CO(2) + NADPH. |
- General function:
- Involved in phosphogluconate dehydratase activity
- Specific function:
- 6-phospho-D-gluconate = 2-dehydro-3-deoxy-6- phospho-D-gluconate + H(2)O
- Gene Name:
- edd
- Uniprot ID:
- P0ADF6
- Molecular weight:
- 64639
Reactions
| 6-phospho-D-gluconate = 2-dehydro-3-deoxy-6-phospho-D-gluconate + H(2)O. |
- General function:
- Involved in shikimate kinase activity
- Specific function:
- ATP + D-gluconate = ADP + 6-phospho-D- gluconate
- Gene Name:
- idnK
- Uniprot ID:
- P39208
- Molecular weight:
- 21004
Reactions
| ATP + D-gluconate = ADP + 6-phospho-D-gluconate. |
- General function:
- Involved in shikimate kinase activity
- Specific function:
- ATP + D-gluconate = ADP + 6-phospho-D- gluconate
- Gene Name:
- gntK
- Uniprot ID:
- P46859
- Molecular weight:
- 19543
Reactions
| ATP + D-gluconate = ADP + 6-phospho-D-gluconate. |
- General function:
- Involved in 6-phosphogluconolactonase activity
- Specific function:
- Catalyzes the hydrolysis of 6-phosphogluconolactone to 6-phosphogluconate
- Gene Name:
- pgl
- Uniprot ID:
- P52697
- Molecular weight:
- 36308
Reactions
| 6-phospho-D-glucono-1,5-lactone + H(2)O = 6-phospho-D-gluconate. |