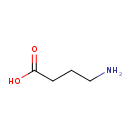
gamma-Aminobutyric acid (ECMDB04017) (M2MDB000561)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2012-05-31 14:02:20 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2015-06-03 15:54:37 -0600 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Name: | gamma-Aminobutyric acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Gamma-aminobutyric acid (GABA) is a metabolite of glutamate. Gamma-aminobutyric acid was first synthesized in 1883, and was first known only as a plant and microbial metabolic product. In 1950, however, GABA was discovered to be an integral part of the mammalian central nervous system. Organisms synthesize GABA from glutamate using the enzyme L-glutamic acid decarboxylase and pyridoxal phosphate (which is the active form of vitamin B6) as a cofactor. (Wikipedia) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula: | C4H9NO2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Weight: | Average: 103.1198 Monoisotopic: 103.063328537 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key: | BTCSSZJGUNDROE-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI: | InChI=1S/C4H9NO2/c5-3-1-2-4(6)7/h1-3,5H2,(H,6,7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS number: | 56-12-2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name: | 4-aminobutanoic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional IUPAC Name: | gamma(amino)-butyric acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES: | NCCCC(O)=O | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as gamma amino acids and derivatives. These are amino acids having a (-NH2) group attached to the gamma carbon atom. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carboxylic acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Amino acids, peptides, and analogues | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Gamma amino acids and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aliphatic acyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State: | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Charge: | 0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Melting point: | 203 °C | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations: | Cytoplasm | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reactions: | gamma-Aminobutyric acid + alpha-Ketoglutarate <> L-Glutamate + Succinic acid semialdehyde L-Glutamate + Hydrogen ion <> gamma-Aminobutyric acid + Carbon dioxide 4-(Glutamylamino) butanoate + Water <> gamma-Aminobutyric acid + L-Glutamate 4-Aminobutyraldehyde + Water + NAD <> gamma-Aminobutyric acid +2 Hydrogen ion + NADH L-Glutamate <> gamma-Aminobutyric acid + Carbon dioxide 4-Aminobutyraldehyde + NAD + Water <> gamma-Aminobutyric acid + NADH + Hydrogen ion 4-Aminobutyraldehyde + NAD + Water > gamma-Aminobutyric acid + NADH + Hydrogen ion Oxoglutaric acid + gamma-Aminobutyric acid <> L-Glutamate + Succinic acid semialdehyde Hydrogen ion + L-Glutamate > Carbon dioxide + gamma-Aminobutyric acid 4-(Glutamylamino) butanoate + Water > gamma-Aminobutyric acid + L-Glutamate 4-Aminobutyraldehyde + NAD + Water > gamma-Aminobutyric acid + NADH L-Glutamate > gamma-Aminobutyric acid + Carbon dioxide gamma-Aminobutyric acid + Oxoglutaric acid > Succinic acid semialdehyde + L-Glutamate 4-(Glutamylamino) butanoate + Water > L-Glutamic acid + gamma-Aminobutyric acid + L-Glutamate gamma-Aminobutyric acid + Oxoglutaric acid > Succinic acid semialdehyde + L-Glutamic acid + L-Glutamate gamma-glutamyl-gamma-aminobutyrate + Water > gamma-Aminobutyric acid + L-Glutamate gamma-Aminobutyric acid + alpha-Ketoglutarate <> L-Glutamate + Succinic acid semialdehyde gamma-Aminobutyric acid + alpha-Ketoglutarate <> L-Glutamate + Succinic acid semialdehyde | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMPDB Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EcoCyc Pathways: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Find out more about how we convert literature concentrations. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra: | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference: | Minoshima, Ryoichi. Preparation of gamma-aminobutyric acid with unripened beans. PCT Int. Appl. (2007), 25pp. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Material Safety Data Sheet (MSDS) | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links: |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in 4-aminobutyrate transaminase activity
- Specific function:
- 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate
- Gene Name:
- gabT
- Uniprot ID:
- P22256
- Molecular weight:
- 45774
Reactions
| 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate. |
| (S)-3-amino-2-methylpropanoate + 2-oxoglutarate = 2-methyl-3-oxopropanoate + L-glutamate. |
- General function:
- Involved in 4-aminobutyrate transaminase activity
- Specific function:
- Involved in the breakdown of putrescine via transamination of gamma-aminobutyrate
- Gene Name:
- puuE
- Uniprot ID:
- P50457
- Molecular weight:
- 44729
Reactions
| 4-aminobutanoate + 2-oxoglutarate = succinate semialdehyde + L-glutamate. |
- General function:
- Involved in glutamate decarboxylase activity
- Specific function:
- Converts glutamate to gamma-aminobutyrate (GABA), consuming one intracellular proton in the reaction. The gad system helps to maintain a near-neutral intracellular pH when cells are exposed to extremely acidic conditions. The ability to survive transit through the acidic conditions of the stomach is essential for successful colonization of the mammalian host by commensal and pathogenic bacteria
- Gene Name:
- gadA
- Uniprot ID:
- P69908
- Molecular weight:
- 52685
Reactions
| L-glutamate = 4-aminobutanoate + CO(2). |
- General function:
- Involved in glutamate decarboxylase activity
- Specific function:
- Converts glutamate to gamma-aminobutyrate (GABA), consuming one intracellular proton in the reaction. The gad system helps to maintain a near-neutral intracellular pH when cells are exposed to extremely acidic conditions. The ability to survive transit through the acidic conditions of the stomach is essential for successful colonization of the mammalian host by commensal and pathogenic bacteria
- Gene Name:
- gadB
- Uniprot ID:
- P69910
- Molecular weight:
- 52668
Reactions
| L-glutamate = 4-aminobutanoate + CO(2). |
- General function:
- Involved in hydrolase activity
- Specific function:
- Involved in the breakdown of putrescine via hydrolysis of the gamma-glutamyl linkage of gamma-glutamyl-gamma- aminobutyrate
- Gene Name:
- puuD
- Uniprot ID:
- P76038
- Molecular weight:
- 28013
Reactions
| 4-(L-gamma-glutamylamino)butanoate + H(2)O = 4-aminobutanoate + L-glutamate. |
- General function:
- Involved in aminobutyraldehyde dehydrogenase activity
- Specific function:
- Catalyzes the oxidation of 1-pyrroline, which is spontaneously formed from 4-aminobutanal, leading to 4- aminobutanoate (GABA). Can also oxidize n-alkyl medium-chain aldehydes, but with a lower catalytic efficiency
- Gene Name:
- ydcW
- Uniprot ID:
- P77674
- Molecular weight:
- 50830
Reactions
| 4-aminobutanal + NAD(+) + H(2)O = 4-aminobutanoate + NADH. |
Transporters
- General function:
- Involved in transport
- Specific function:
- Transporter for GABA
- Gene Name:
- gabP
- Uniprot ID:
- P25527
- Molecular weight:
- 51080
- General function:
- Involved in amino acid transmembrane transporter activity
- Specific function:
- Involved in glutamate-dependent acid resistance. Imports glutamate inside the cell while simultaneously exporting to the periplasm the GABA produced by gadA and gadB. The gad system helps to maintain a near-neutral intracellular pH when cells are exposed to extremely acidic conditions. The ability to survive transit through the acidic conditions of the stomach is essential for successful colonization of the mammalian host by commensal and pathogenic bacteria
- Gene Name:
- gadC
- Uniprot ID:
- P63235
- Molecular weight:
- 55076
- General function:
- Involved in transporter activity
- Specific function:
- Non-specific porin
- Gene Name:
- ompN
- Uniprot ID:
- P77747
- Molecular weight:
- 41220
- General function:
- Involved in transporter activity
- Specific function:
- Uptake of inorganic phosphate, phosphorylated compounds, and some other negatively charged solutes
- Gene Name:
- phoE
- Uniprot ID:
- P02932
- Molecular weight:
- 38922
- General function:
- Involved in transporter activity
- Specific function:
- OmpF is a porin that forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane. It is also a receptor for the bacteriophage T2
- Gene Name:
- ompF
- Uniprot ID:
- P02931
- Molecular weight:
- 39333
- General function:
- Involved in transporter activity
- Specific function:
- Forms passive diffusion pores which allow small molecular weight hydrophilic materials across the outer membrane
- Gene Name:
- ompC
- Uniprot ID:
- P06996
- Molecular weight:
- 40368